


Design-adjusted inference for pathogen lineage surveillance
under
unequal sequencing and reporting delays
Genomic surveillance systems sequence unevenly. Denmark sequences 12% of cases; Romania sequences 0.3%. If you estimate lineage prevalence by counting sequences, the result is dominated by Denmark — regardless of what is actually circulating across Europe.
On real ECDC data, this produces up to 14 percentage points of error:

The red shaded area is the bias eliminated by design weighting. survinger corrects this using Horvitz-Thompson / Hajek estimators with Wilson score confidence intervals.

Each country’s bias depends on its sequencing rate and its local prevalence, and both change over time. Poland (under-sequenced, high prevalence) is systematically underweighted by naive methods. A single correction factor cannot fix this — you need per-stratum, per-period weights.
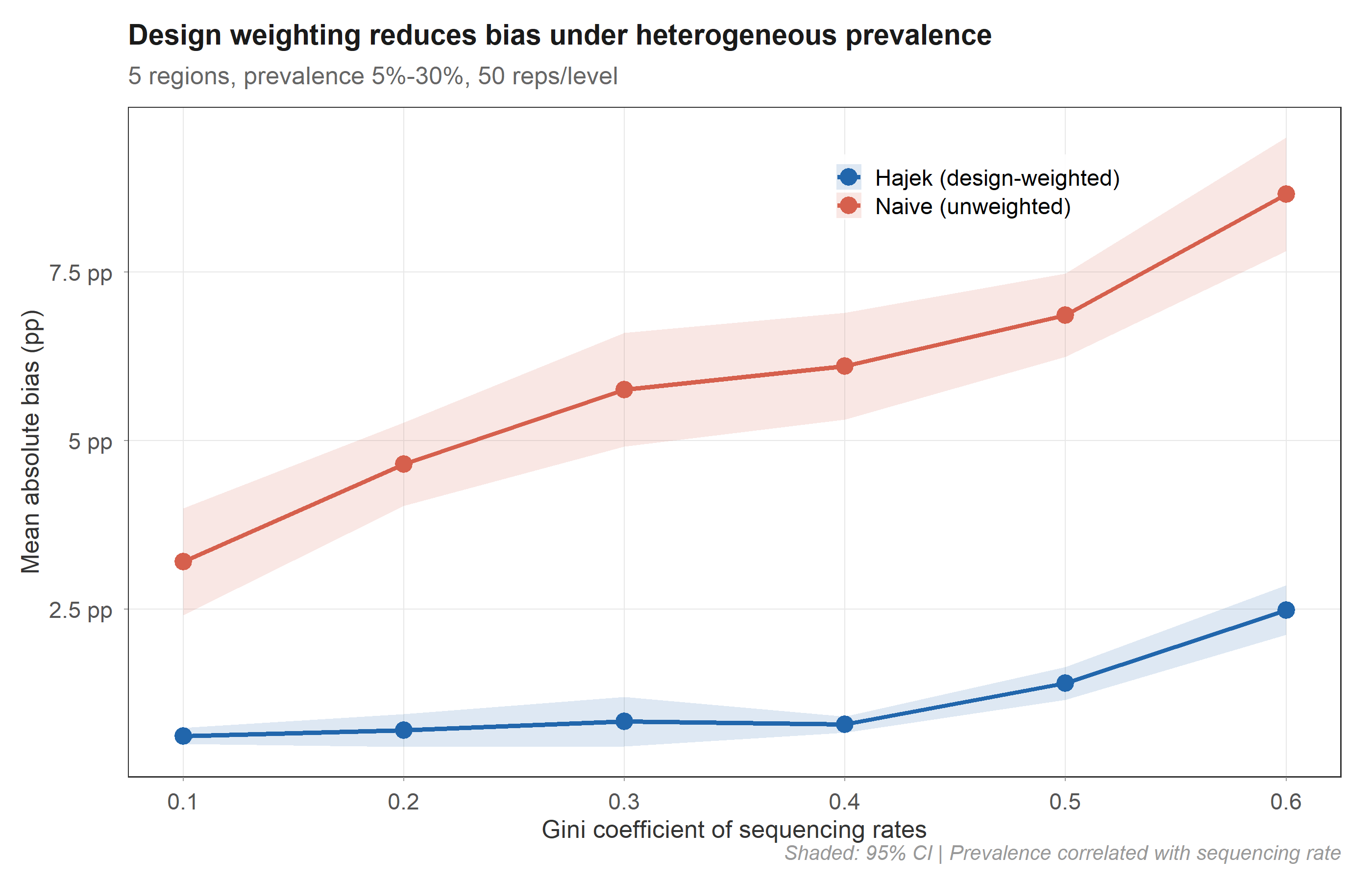
In controlled simulation (50 replicates × 6 inequality levels), the Hajek estimator maintains 0.6–2.5 pp absolute bias while the naive estimator reaches 3.2–8.7 pp. The advantage holds across all levels of sequencing inequality.
# install.packages("remotes")
remotes::install_github("CuiweiG/survinger")library(survinger)
# Simulate surveillance data (or use your own)
sim <- surv_simulate(n_regions = 5, n_weeks = 26, seed = 42)
# Create design from surveillance data
design <- surv_design(
data = sim$sequences, strata = ~ region,
sequencing_rate = sim$population[c("region", "seq_rate")],
population = sim$population
)
# Corrected prevalence (one line)
surv_lineage_prevalence(design, "BA.2.86")
# Or even simpler — single pipe-friendly call:
surv_estimate(
data = sim$sequences, strata = ~ region,
sequencing_rate = sim$population[c("region", "seq_rate")],
population = sim$population, lineage = "BA.2.86"
)
# Full pipeline with delay correction
delay <- surv_estimate_delay(design)
surv_adjusted_prevalence(design, delay, "BA.2.86")
# How should I allocate 500 sequences?
surv_optimize_allocation(design, "min_mse", total_capacity = 500)
# Is my system powerful enough?
surv_detection_probability(design, true_prevalence = 0.01)
# One-page diagnostic
surv_report(design)| Function | Purpose |
|---|---|
surv_design() |
Create design with inverse-probability weights |
surv_simulate() |
Generate synthetic surveillance data |
surv_filter() |
Subset a design by filter criteria |
surv_update_rates() |
Update sequencing rates |
surv_set_weights() |
Override design weights |
| Function | Purpose |
|---|---|
surv_lineage_prevalence() |
Hajek / HT / post-stratified prevalence |
surv_naive_prevalence() |
Unweighted baseline prevalence |
surv_prevalence_by() |
Prevalence by subgroup (region, source, etc.) |
surv_estimate() |
Pipe-friendly one-call analysis |
| Function | Purpose |
|---|---|
surv_estimate_delay() |
Right-truncation-corrected delay fitting |
surv_reporting_probability() |
Cumulative reporting probability |
surv_nowcast_lineage() |
Delay-adjusted nowcast |
surv_adjusted_prevalence() |
Combined design + delay correction |
| Function | Purpose |
|---|---|
surv_optimize_allocation() |
Neyman allocation (3 objectives) |
surv_compare_allocations() |
Benchmark all allocation strategies |
surv_required_sequences() |
Sample size for target detection power |
| Function | Purpose |
|---|---|
surv_detection_probability() |
Variant detection power |
surv_power_curve() |
Detection probability across prevalence range |
surv_compare_estimates() |
Weighted vs naive side-by-side plot |
surv_design_effect() |
Design effect over time |
surv_sensitivity() |
Sensitivity analysis across all methods |
surv_report() |
Surveillance system diagnostic |
surv_quality() |
One-row quality metrics |
| Function | Purpose |
|---|---|
tidy() / glance() |
Broom-style tidying for all result objects |
surv_bind() |
Combine multiple prevalence estimates |
surv_table() |
Publication-ready formatted table |
theme_survinger() |
Publication-quality ggplot2 theme |
| phylosamp | survey | epinowcast | survinger | |
|---|---|---|---|---|
| Question | How many? | General surveys | Bayesian nowcast | Allocate + correct + nowcast |
| Genomic-specific | ✓ | ✗ | Partial | ✓ |
| Allocation | ✗ | ✗ | ✗ | ✓ (3 objectives) |
| Delay correction | ✗ | ✗ | ✓ | ✓ |
| Requires Stan | ✗ | ✗ | ✓ | ✗ |
| CRAN-friendly | ✓ | ✓ | ✗ | ✓ |
survey::svymean (exact
match)vignette("survinger") — Quick startvignette("allocation-optimization") — Resource
allocationvignette("delay-correction") — Delay estimation and
nowcastingvignette("real-world-ecdc") — ECDC case study@Manual{survinger2026,
title = {survinger: Design-Adjusted Inference for Pathogen Lineage Surveillance},
author = {Cuiwei Gao},
year = {2026},
note = {R package version 0.1.0},
url = {https://github.com/CuiweiG/survinger}
}MIT © 2026 Cuiwei Gao